Dr. Chinyere Ekine-Dzivenu

Picture 1. Training Cohort and Faculty: Participants and trainers at the integrated training on livestock genetics, genomics, and leadership held at ILRI, Nairobi (16–27 February 2026).
As part of my joint appointment, with 20% effort dedicated to the International Livestock Research Institute (ILRI), I participated in and contributed to an integrated training program on Genetic and Genomic Approaches for Livestock Improvement and a leadership-focused Training on Enhancing Capacity and Skills for Livestock Impact, held at ILRI, Nairobi, Kenya, from 16–27 February 2026. This program combined advanced technical training in animal genetics and genomics with leadership development, strengthening capacity to design and implement sustainable breeding programs and support livestock system transformation in the global south. The training was delivered by an international faculty drawn from leading research institutions, including the University of Alberta, the University of Edinburgh, ILRI, the Centre for Tropical Livestock Genetics and Health (CTLGH), and Scotland’s Rural College (SRUC), reflecting a high level of scientific expertise and global collaboration.
The technical component provided both theoretical and hands-on training in key areas of modern animal breeding, including phenotypic data analysis and linear models in R, pedigree-based genetic evaluation using BLUP, genomic data analysis with SNPs etc. I contributed through lectures, facilitation, and mentorship. I also moderated part of the African Animal Breeders Network (AABNET) book launch on African livestock genetic resources. which marked the release of “African Livestock Genetic Resources and Sustainable Breeding Strategies: Unlocking a Treasure Trove”.

Picture 2. ILRI Leadership at Book Launch: The Director General of ILRI Appolinaire Djikeng holding the book African Livestock Genetic Resources and Sustainable Breeding Strategies: Unlocking a Treasure Trove during the AABNet book launch.
This book is the first to comprehensively present, in a single source, the diversity and uniqueness of African livestock genetic resources alongside sustainable breeding strategies and emerging technologies. The program engaged a diverse international cohort, with participants from over 25 countries across Africa and Asia representing academia, research institutions, government, and development organizations, thereby contributing directly to capacity building and livestock development in low- and middle-income countries.
Overall, the integrated training strengthened technical expertise, leadership and mentoring capacity, and international collaboration, while reinforcing partnerships between ILRI and the University of Alberta and Livestock Gentec. This engagement contributes to advancing livestock development in low- and middle-income countries through improved breeding strategies and knowledge transfer, while also enhancing the global research profile, partnerships, and impact of the University of Alberta and Livestock Gentec in the field of livestock genetics and sustainable agricultural development.


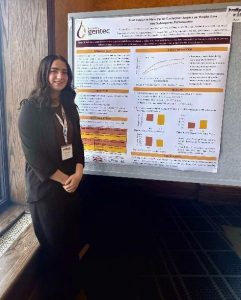
 Sonja Allen (PhD candidate)
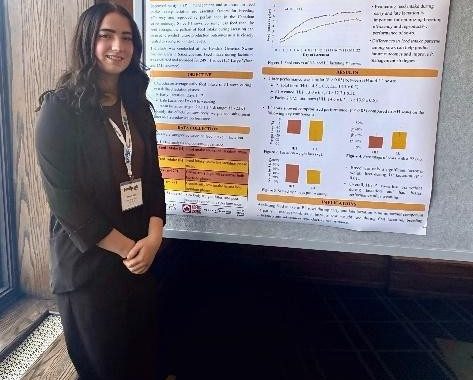
Sonja Allen (PhD candidate) Kayla Patey (MSc candidate)
Kayla Patey (MSc candidate)